There are two types of acetylation processes widely occurred in proteins. The first Nα-terminal acetylation is catalyzed a variety of N-terminal acetyltransferases (NATs), which cotranslationally transfer acetyl moieties from acetyl-coenzyme A (Acetyl-CoA) to the α-amino (Nα) group of protein amino-terminal residues. Although Nα-terminal acetylation is rare in prokaryotes, it was estimated that about 85% of eukaryotic proteins are Nα-terminally modified (Polevoda et al., 2000; Polevoda et al., 2002). The second type is Nε-lysine acetylation, which specifically modifies ε-amino group of protein lysine residues (Yang et al., 2007; Shahbazian et al., 2007; Smith et al., 2009). Although Nε-lysine acetylation is less common, it's one of the most important and ubiquitous post-translational modifications conserved in prokaryotes and eukaryotes. Moreover, the acetylation and deacetylation are dynamically and temporally regulated by histone acetyltransferases (HATs) and histone deacetylases (HDACs), respectively (Yang et al., 2004; Lee et al., 2007).
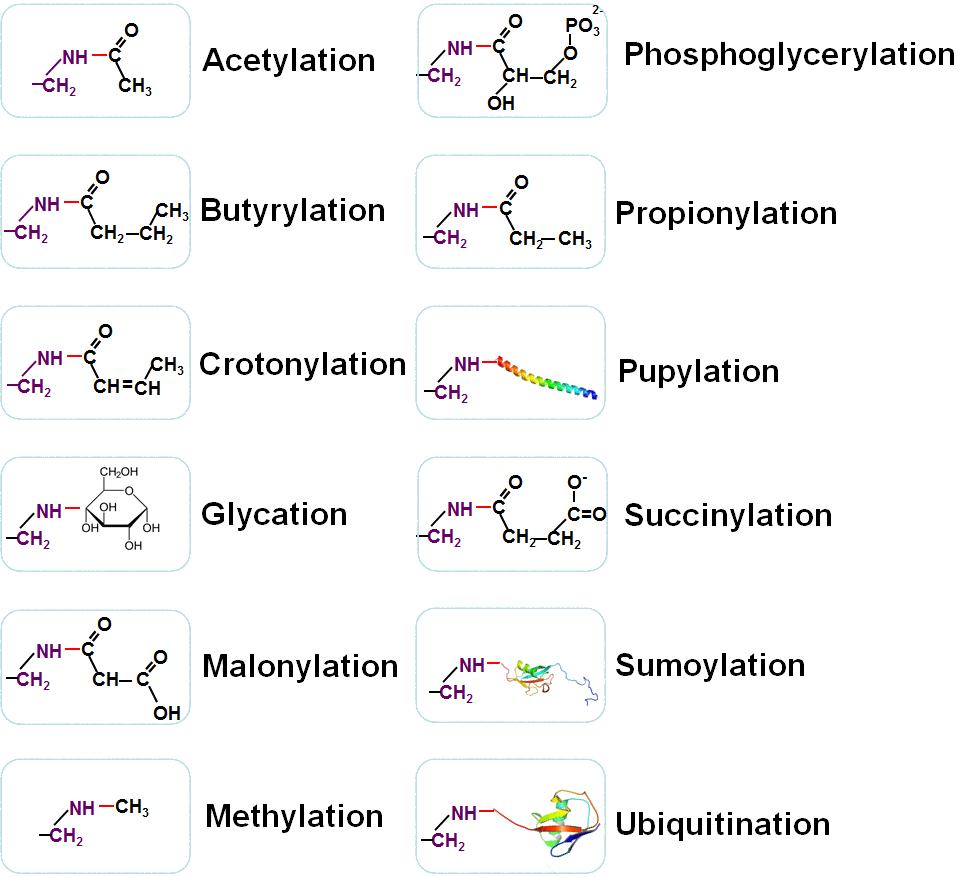
Beyond lysine acetylation, there are also a dozen of other types of post-translational modifications (PTMs) extensively occurred at lysine residues, such as ubiquitination (Gao, et al., 2013), methylation (Chen, et al., 2006), sumoylation (Ren, et al., 2009; Xue, et al., 2006), glycation (Priego-Capote, et al., 2010), butyrylation (Chen, et al., 2007; Cheng, et al., 2009; Zhang, et al., 2009), crotonylation (Tan, et al., 2011), malonylation (Xie, et al., 2012), propionylation (Chen, et al., 2007; Cheng, et al., 2009; Zhang, et al., 2009), succinylation (Xie, et al., 2012; Zhang, et al., 2011), phosphoglycerylation (Moellering, R. E. and B. F. Cravatt, 2013) and prokaryotic Pupylation (Liu, et al., 2011). Thus, the lysine residue can be a "hotspot" of the PTM crosstalk (Lu, et al., 2011; Minguez, et al., 2012; van Noort, et al., 2012). Identification of these well-known or novel lysine modifications is fundamental for understanding their molecular mechanisms and regulatory roles.
Thus, we developed an integrative database of CPLM (Compendium of Protein Lysine Modifications) for 12 types of protein lysine modifications, including 203,972 modification events on 189,919 modified lysines in 45,748 proteins. Because this database was adapted and extended from our CPLA (Compendium of Protein Lysine Acetylation) 1.0 database, which was lastly updated on March 1st, 2010, containing 3,311 unique protein entries with 7,151 lysine acetylation sites, the current version of CPLM is 2.0. The database can be browse by either 1) types or 2) species.
| Browse by types | Browse by species |
|

|



